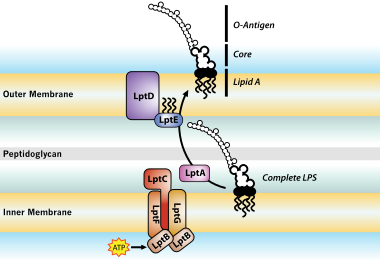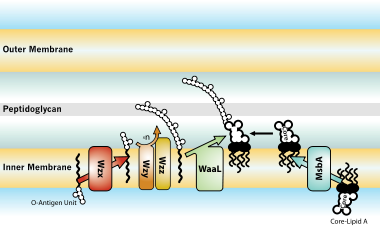
Transport of lipopolysaccharide across the cell envelope: the long road of discovery | Nature Reviews Microbiology
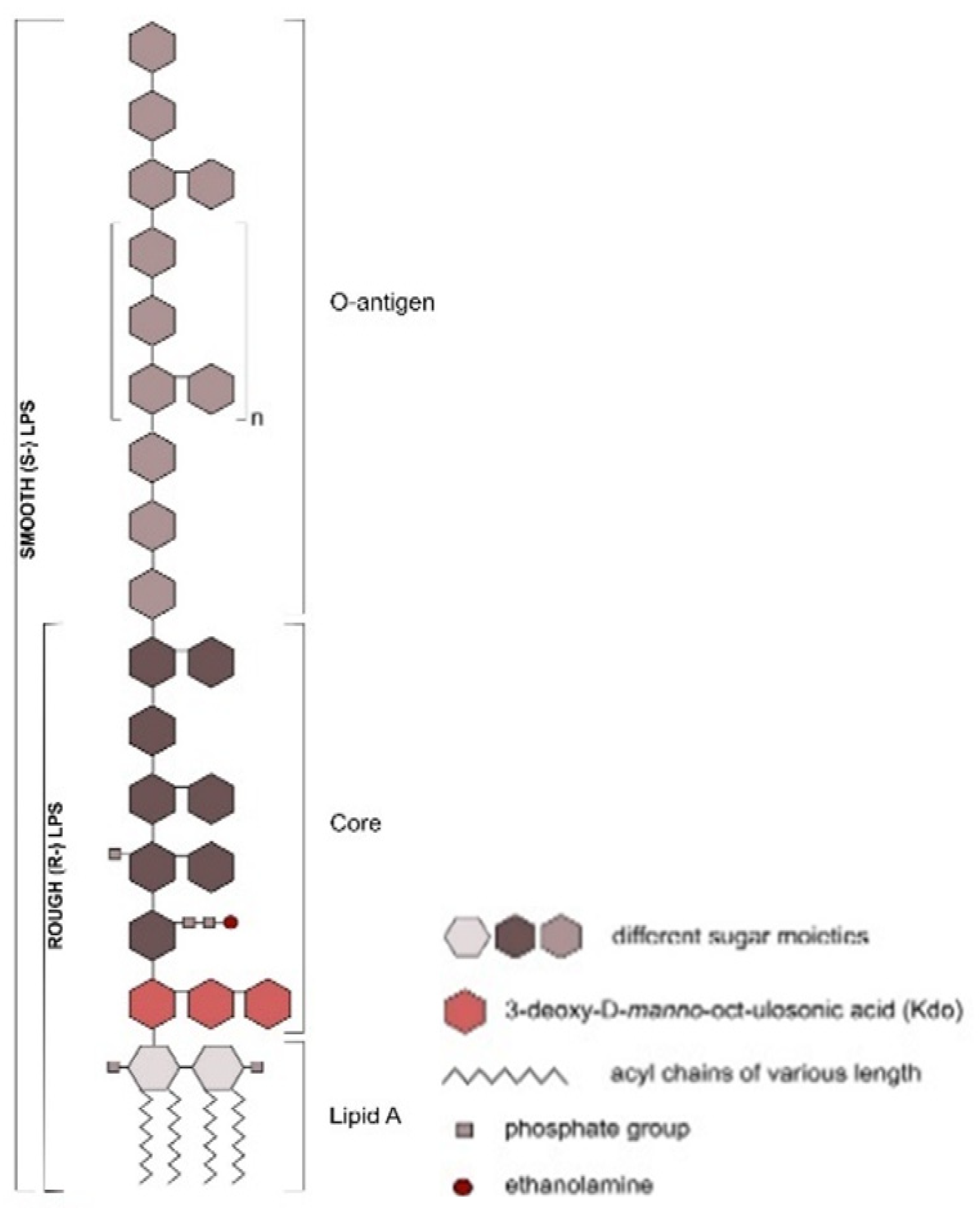
Animals | Free Full-Text | Ruminal Lipopolysaccharides Analysis: Uncharted Waters with Promising Signs

LPS and O-antigen synthesis. (A) Model representing LPS and O-antigen... | Download Scientific Diagram

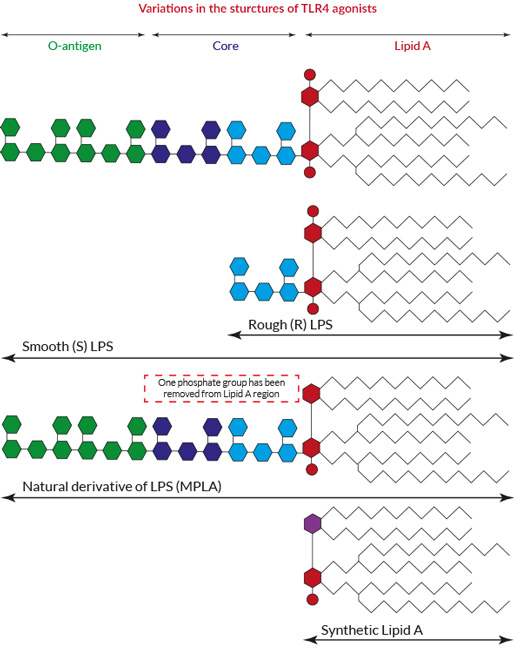
The Role of Lipopolysaccharide-Induced Cell Signalling in Chronic Inflammation - Martin J. Page, Douglas B. Kell, Etheresia Pretorius, 2022

Fighting Against Bacterial Lipopolysaccharide-Caused Infections through Molecular Dynamics Simulations: A Review | Journal of Chemical Information and Modeling
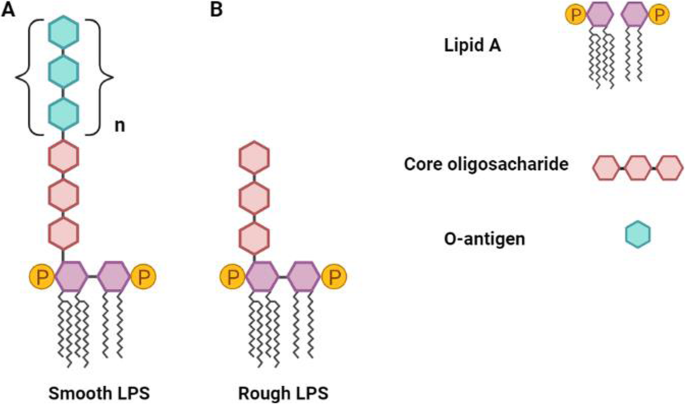
Ruminal bacteria lipopolysaccharides: an immunological and microbial outlook | Journal of Animal Science and Biotechnology | Full Text

Pathogens | Free Full-Text | The Role of Pseudomonas aeruginosa Lipopolysaccharide in Bacterial Pathogenesis and Physiology
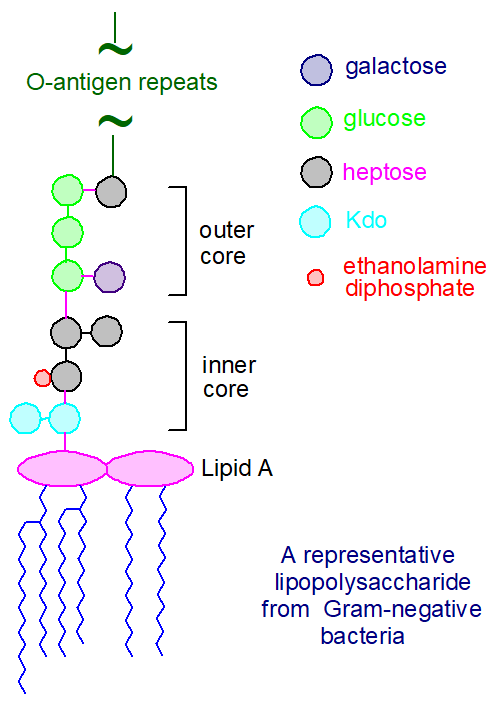
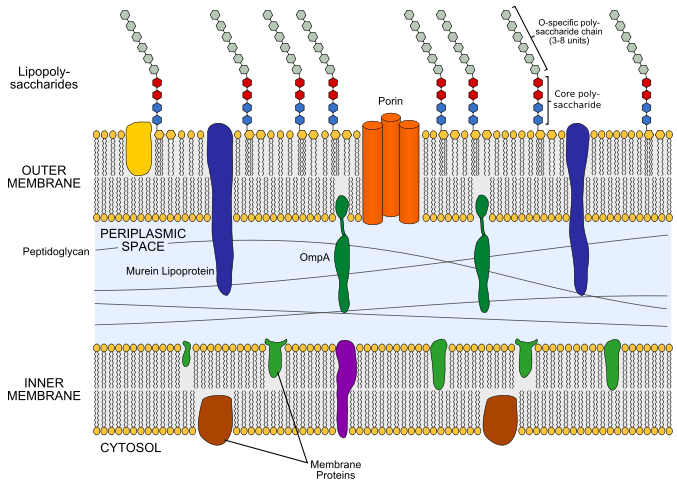
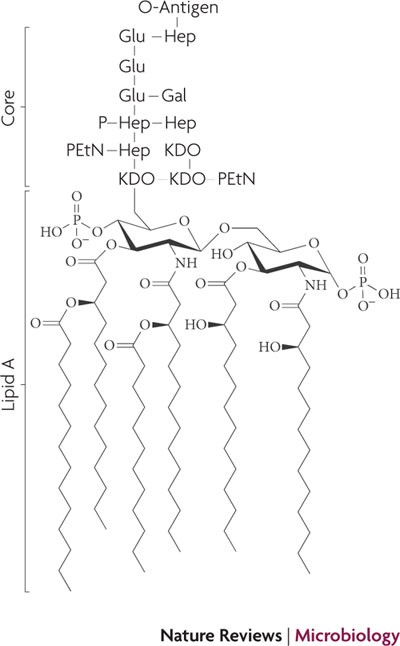
Lipid A and Bacterial Lipopolysaccharides, Gram-negative, endotoxins, lipo-chitooligosaccharides (Nod factors) - structure, occurrence, biochemistry and function

Lipopolysaccharide O-antigen molecular and supramolecular modifications of plant root microbiota are pivotal for host recognition - ScienceDirect

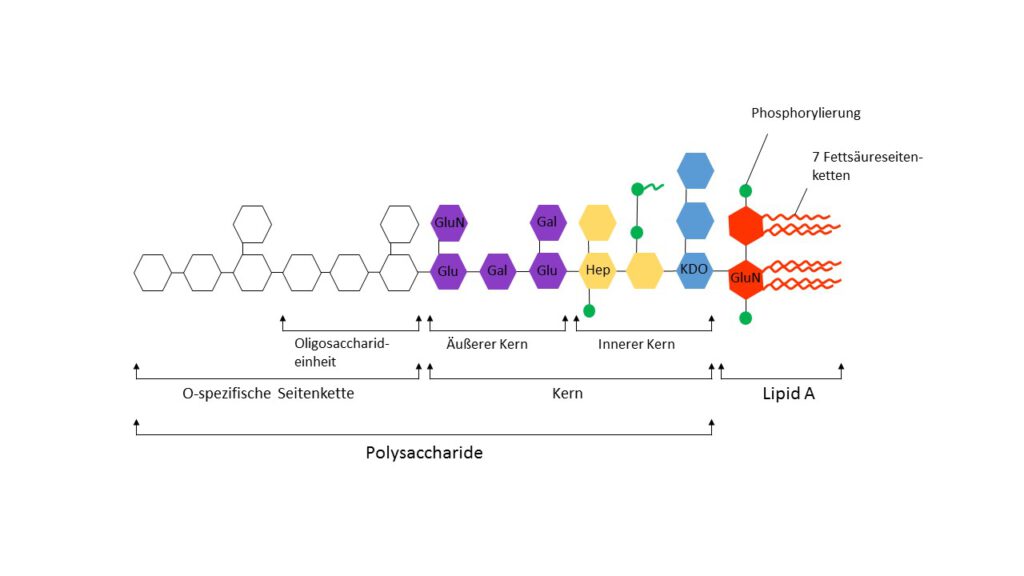
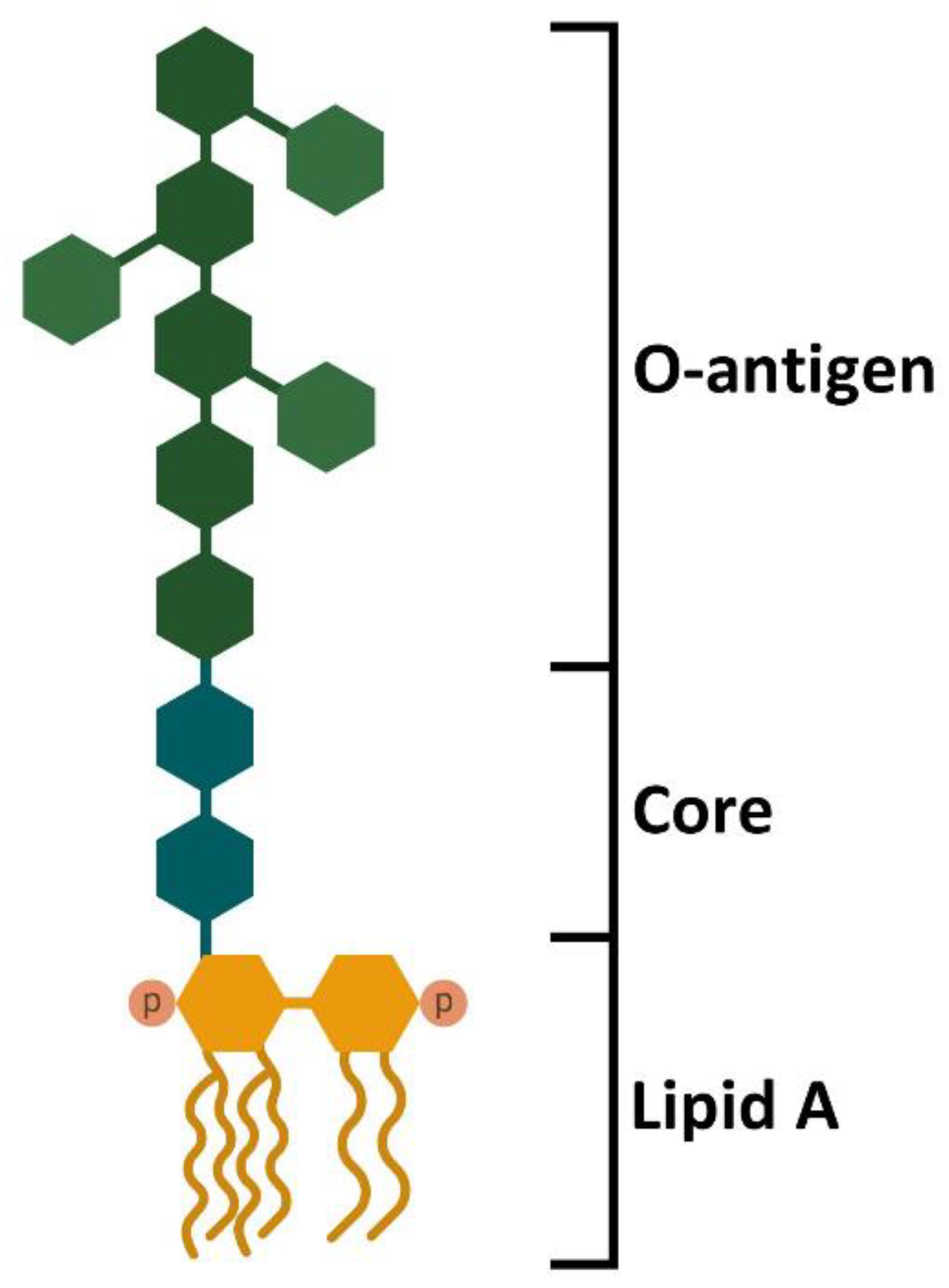
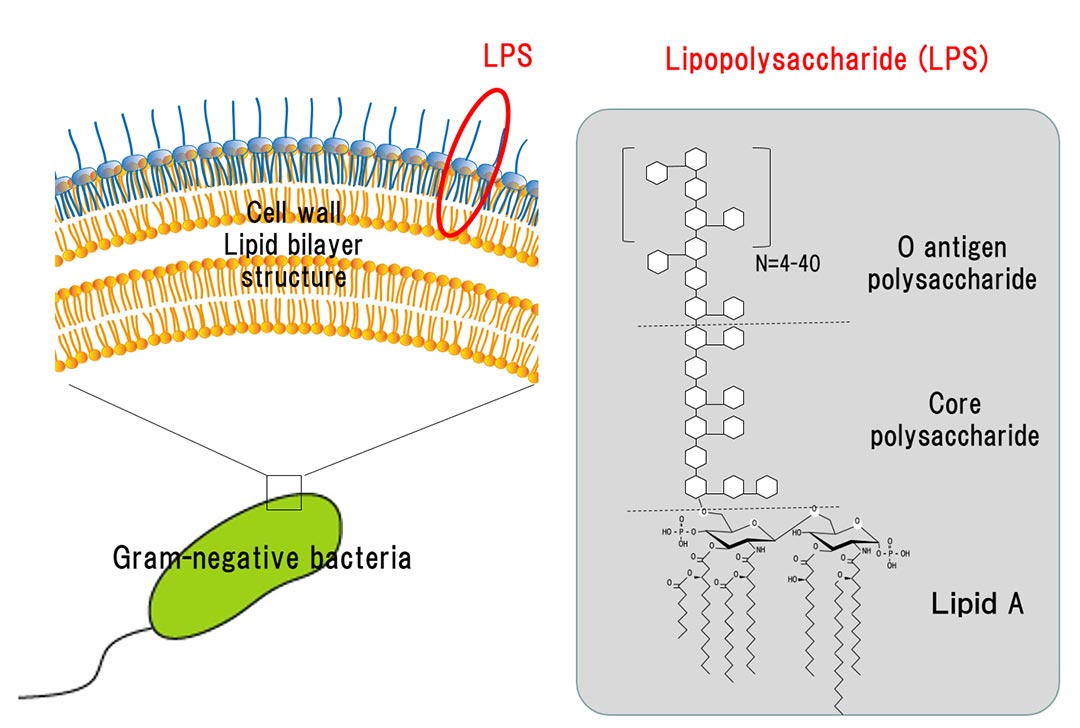
Schematic of the basic structure of lipopolysaccharide. LPS consists of... | Download Scientific Diagram
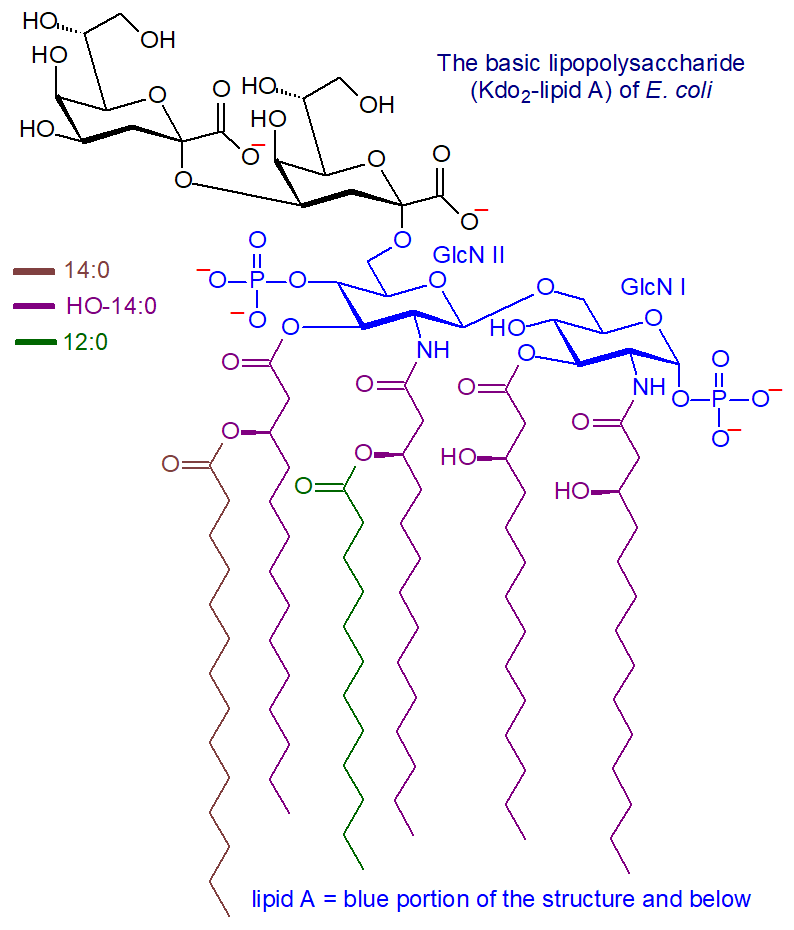
Lipid A and Bacterial Lipopolysaccharides, Gram-negative, endotoxins, lipo-chitooligosaccharides (Nod factors) - structure, occurrence, biochemistry and function